With the inlib/wroot classes you can write histograms and ntuples in a file at the CERN-ROOT format. Here an "ntuple" is understood as a "flat TTree". With inlib/rroot classes you can read this kind of file. And if the zip compression is used, the exlib/rroot code permits to read "zip compressed" files. You can have a writing example with :
cd <path to inlib>
cd examples/cpp
c++ -I../.. wroot.cpp
./a.out
The program produces the wroot.root file that you can read with :
<source setup CERN-ROOT> # in order to have root in your PATH.
root ../../../exlib/examples/cern_root/rroot.C
You should see something as :
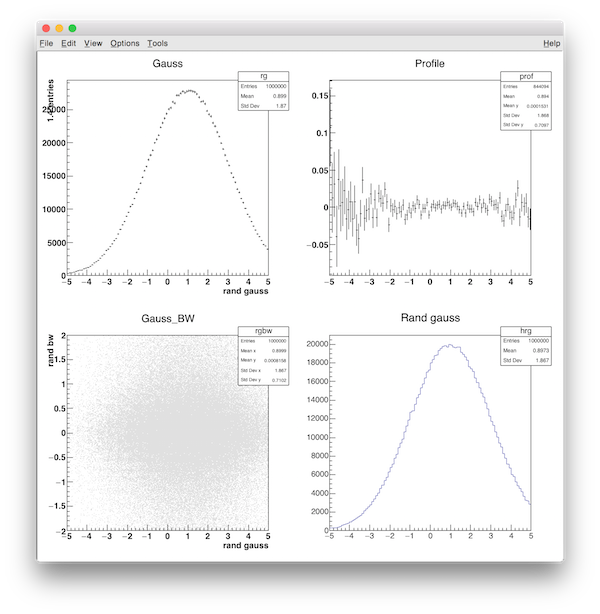
You can also dump the list of "TKeys" of wroot.root by using the exlib/example/cpp/rroot.cpp program :
cd <path to exlib>
cd examples/cpp
./build rroot.cpp
Linux> ./bin_gnu/rroot -ls ../../../inlib/examples/cpp/wroot.root
You should see something as :
format version 40000 object : "rg_rbw", class : TTree object : "rg_rbw_2", class : TTree object : "empty", class : TTree directory : histo object : "rg", class : TH1D object : "rf", class : TH1F object : "prof", class : TProfile
The rpawdemo.cpp program
The rpawdemo.cpp program permits to read and dump an histo from the pawdemo.root file that comes from the distribution.
cd <path to exlib>
cd examples/cpp
./build rpawdemo.cpp
cp ../../data/pawdemo.root .
Linux> ./bin_gnu/rpawdemo
( Darwin> ./bin_clan/rpawdemo )
( cygwin> ./bin_visual/rpawdemo.exe )
You should see something as :
size of h10 : 900 1TEST1 * ENTRIES = 10000 * ALL CHANNELS = 7314 * UNDERFLOW = 14 * OVERFLOW = 14 * BIN WID = 0.06 * MEAN VALUE = 0.00538966 * R . M . S = 1.1085 * ENTRIES[0] = 0 * HEIGHT[0] = 3 * ERROR[0] = 0 * ENTRIES[N/2] = 0 * HEIGHT[N/2] = 127 * ERROR[N/2] = 0 * ENTRIES[N-1] = 0 * HEIGHT[N-1] = 1 * ERROR[N-1] = 0
// Copyright (C) 2010, Guy Barrand. All rights reserved.
// See the file exlib.license for terms.
//exlib_build_use exlib inlib csz
#include <inlib/mem>
#include <inlib/args>
#include <inlib/rroot/file>
#include <inlib/rroot/streamers>
#include <iostream>
#include <cstdlib>
int main(int argc,char** argv) {
#ifdef INLIB_MEM
inlib::mem::set_check_by_class(true);{
#endif
inlib::args args(argc,argv);
bool verbose = args.is_arg("-verbose");
std::string file = "pawdemo.root";
inlib::rroot::file rfile(std::cout,file,verbose);
inlib::rroot::key* key = rfile.dir().find_key("h10");
if(!key) {
std::cout << "key for h10 not found." << std::endl;
return EXIT_FAILURE;
}
unsigned int sz;
char* buf = key->get_object_buffer(rfile,sz);
if(!buf) {
std::cout << "can't get data buffer for h10." << std::endl;
return EXIT_FAILURE;
}
std::cout << "size of h10 : " << sz << std::endl;
inlib::rroot::buffer b(std::cout,rfile.byte_swap(),sz,buf,key->key_length(),verbose);
inlib::histo::h1d* h = inlib::rroot::TH1F_stream(b); //we get ownership on h.
if(!h) {
std::cout << "streaming failed for h10." << std::endl;
return EXIT_FAILURE;
}
h->hprint(std::cout);
delete h;
#ifdef INLIB_MEM
}inlib::mem::balance(std::cout);
#endif
return EXIT_SUCCESS;
}
The wroot.cpp program
// Copyright (C) 2010, Guy Barrand. All rights reserved.
// See the file inlib.license for terms.
// This program produces a wroot.root file.
//
// See rroot.C for an example of how to manipulate
// (and check !) the content of this file with CERN-ROOT.
//inlib_build_use kernel
#ifndef EXLIB_DONT_HAVE_ZLIB
#define EXLIB_DONT_HAVE_ZLIB
#endif
#ifdef INLIB_MEM
#include <inlib/mem>
#endif //INLIB_MEM
#include <inlib/wroot/file>
#include <inlib/wroot/to>
#include <inlib/histo/h1d>
#include <inlib/histo/h2d>
#include <inlib/histo/h3d>
#include <inlib/histo/p1d>
#include <inlib/histo/p2d>
#include <inlib/wroot/ntuple>
#include <inlib/histo/h1df>
#include <inlib/randd>
#include <inlib/randf>
#include <inlib/rroot/file>
#include <inlib/rroot/ntuple>
#include <inlib/rroot/fac>
#include <inlib/eqT>
#ifdef EXLIB_DONT_HAVE_ZLIB
#else
#include <exlib/zlib>
#endif
#include <inlib/args>
#include <iostream>
#include <cstdlib>
int main(int argc,char** argv) {
#ifdef INLIB_MEM
inlib::mem::set_check_by_class(true);{
#endif //INLIB_MEM
inlib::args args(argc,argv);
bool verbose = args.is_arg("-verbose");
bool row_wise = args.is_arg("-row_wise");
//////////////////////////////////////////////////////////
/// create a .root file : ////////////////////////////////
//////////////////////////////////////////////////////////
if(verbose) std::cout << (row_wise?"row_wise":"column_wise") << std::endl;
std::string file = "wroot.root";
{inlib::wroot::file rfile(std::cout,file);
#ifdef EXLIB_DONT_HAVE_ZLIB
#else
if(args.is_arg("-noz")){
} else {
rfile.add_ziper('Z',exlib::compress_buffer);
rfile.set_compression(9);
}
#endif
bool osc_stream = args.is_arg("-osc");
inlib::wroot::directory* dir = rfile.dir().mkdir("histo");
if(!dir) {
std::cout << "can't create diectory." << std::endl;
return EXIT_FAILURE;
}
//inlib::wroot::directory* dxxx = dir->mkdir("xxx");
//if(!dxxx) {
// std::cout << "can't create diectory." << std::endl;
// return EXIT_FAILURE;
//}
//////////////////////////////////////////////////////////
/// create some histos : /////////////////////////////////
//////////////////////////////////////////////////////////
unsigned int num_megas;
args.find<unsigned int>("-megas",num_megas,1);
unsigned int entries = num_megas*1000000;
inlib::rgaussd rg(1,2);
inlib::rbwd rbw(0,1);
{inlib::histo::h1d h("Gauss",100,-5,5);
for(unsigned int count=0;count<entries;count++) h.fill(rg.shoot(),1.4);
// plotting hints :
h.add_annotation(inlib::histo::key_axis_x_title(),"rand gauss");
h.add_annotation(inlib::histo::key_axis_y_title(),"1.4*entries");
if(verbose) {
std::cout << "h1d : " << h.title()
<< ", all entries " << h.all_entries()
<< ", entries " << h.entries()
<< ", mean " << h.mean() << ", rms " << h.rms()
<< std::endl;
}
// write :
if(osc_stream) {
if(!inlib::wroot::to_osc(*dir,h,"rg")) return EXIT_FAILURE;
} else {
if(!inlib::wroot::to(*dir,h,"rg")) return EXIT_FAILURE;
}}
{inlib::rgaussf rf(1,2);
inlib::histo::h1df h("GaussF",100,-5,5);
for(unsigned int count=0;count<entries;count++) h.fill(rf.shoot(),1.4f);
// plotting hints :
h.add_annotation(inlib::histo::key_axis_x_title(),"rand gauss");
h.add_annotation(inlib::histo::key_axis_y_title(),"1.4*entries");
if(verbose) {
std::cout << "h1df : " << h.title()
<< ", all entries " << h.all_entries()
<< ", entries " << h.entries()
<< ", mean " << h.mean() << ", rms " << h.rms()
<< std::endl;
}
// write :
if(!inlib::wroot::to(*dir,h,"rf")) return EXIT_FAILURE;}
{inlib::histo::p1d h("Profile",100,-5,5,-2,2);
for(unsigned int count=0;count<entries;count++) h.fill(rg.shoot(),rbw.shoot(),1);
if(verbose) {
std::cout << "p1d : " << h.title()
<< ", all entries " << h.all_entries()
<< ", entries " << h.entries()
<< ", mean " << h.mean() << ", rms " << h.rms()
<< std::endl;
}
if(osc_stream) {
if(!inlib::wroot::to_osc(*dir,h,"prof")) return EXIT_FAILURE;
} else {
if(!inlib::wroot::to(*dir,h,"prof")) return EXIT_FAILURE;
}}
{inlib::histo::h2d h("Gauss_BW",20,-5,5,20,-2,2);
for(unsigned int count=0;count<entries;count++) h.fill(rg.shoot(),rbw.shoot(),0.8);
//plotting hints :
h.add_annotation(inlib::histo::key_axis_x_title(),"rand gauss");
h.add_annotation(inlib::histo::key_axis_y_title(),"rand bw");
h.add_annotation(inlib::histo::key_axis_z_title(),"0.8*entries");
if(verbose) {
std::cout << "h2d : " << h.title()
<< ", all entries " << h.all_entries()
<< ", entries " << h.entries()
<< ", mean_x " << h.mean_x() << ", rms_x " << h.rms_x()
<< ", mean_y " << h.mean_y() << ", rms_y " << h.rms_y()
<< std::endl;
}
// write :
if(osc_stream) {
if(!inlib::wroot::to_osc(*dir,h,"rgbw")) return EXIT_FAILURE;
} else {
if(!inlib::wroot::to(*dir,h,"rgbw")) return EXIT_FAILURE;
}}
{inlib::histo::p2d h("Profile2D",100,-5,5,100,-5,5,-2,2);
for(unsigned int count=0;count<entries;count++) h.fill(rg.shoot(),rg.shoot(),rbw.shoot(),1);
if(verbose) {
std::cout << "p2d : " << h.title()
<< ", all entries " << h.all_entries()
<< ", entries " << h.entries()
<< ", mean_x " << h.mean_x() << ", rms_x " << h.rms_x()
<< ", mean_y " << h.mean_y() << ", rms_y " << h.rms_y()
<< std::endl;
}
if(osc_stream) {
if(!inlib::wroot::to_osc(*dir,h,"prof2D")) return EXIT_FAILURE;
} else {
if(!inlib::wroot::to(*dir,h,"prof2D")) return EXIT_FAILURE;
}}
{inlib::histo::h3d h("Gauss_Gauss_BW",20,-5,5,20,-5,5,20,-2,2);
for(unsigned int count=0;count<entries;count++) h.fill(rg.shoot(),rg.shoot(),rbw.shoot(),0.8);
//plotting hints :
h.add_annotation(inlib::histo::key_axis_x_title(),"rand gauss");
h.add_annotation(inlib::histo::key_axis_y_title(),"rand gauss");
h.add_annotation(inlib::histo::key_axis_z_title(),"rand bw");
if(verbose) {
std::cout << "h3d : " << h.title()
<< ", all entries " << h.all_entries()
<< ", entries " << h.entries()
<< ", mean_x " << h.mean_x() << ", rms_x " << h.rms_x()
<< ", mean_y " << h.mean_y() << ", rms_y " << h.rms_y()
<< ", mean_z " << h.mean_z() << ", rms_z " << h.rms_z()
<< std::endl;
}
// write :
if(osc_stream) {
if(!inlib::wroot::to_osc(*dir,h,"rggbw")) return EXIT_FAILURE;
} else {
if(!inlib::wroot::to(*dir,h,"rggbw")) return EXIT_FAILURE;
}}
//////////////////////////////////////////////////////////
/// create and fill a ntuple : ///////////////////////////
//////////////////////////////////////////////////////////
{//WARNING : the ntuple can't be on the stack. It is owned by the directory.
inlib::wroot::ntuple* ntu = new inlib::wroot::ntuple(rfile.dir(),"rg_rbw","Randoms",row_wise);
inlib::wroot::ntuple::column<int>* col_count = ntu->create_column<int>("count");
inlib::wroot::ntuple::column<double>* col_rgauss = ntu->create_column<double>("rgauss");
inlib::wroot::ntuple::column<float>* col_rbw = ntu->create_column<float>("rbw");
inlib::wroot::ntuple::column_string* col_str = ntu->create_column_string("string");
std::vector<float> user_vec_f;
ntu->create_column_vector_ref<float>("vec_float",user_vec_f); //pass the ref of user_vec_f.
inlib::wroot::ntuple::std_vector_column<double>* col_vec_d = ntu->create_column_vector<double>("vec_d");
std::vector<std::string> user_vec_s;
ntu->create_column_vector_string_ref("vec_s",user_vec_s,'\n');
if(args.is_arg("-large")){
entries = 300000000; //to test >2Gbytes file.
ntu->set_basket_size(1000000);
}
inlib::rtausmed rflat;
unsigned int vec_float_count = 0;
unsigned int vec_double_count = 0;
inlib::rbwf rbwf(0,1);
std::string stmp;
for(unsigned int count=0;count<entries;count++) {
if(!col_count->fill(count)) {
std::cout << "col_count fill failed." << std::endl;
break;
}
if(!col_rgauss->fill(rg.shoot())) {
std::cout << "col_rgauss fill failed." << std::endl;
break;
}
if(!col_rbw->fill(rbwf.shoot())) {
std::cout << "col_rbw fill failed." << std::endl;
break;
}
if(!inlib::num2s(count,stmp)){}
if(!col_str->fill("str "+stmp)) {
std::cout << "col_str fill failed." << std::endl;
break;
}
{user_vec_f.clear();
unsigned int number = (unsigned int)(10*rflat.shoot());
for(unsigned int i=0;i<number;i++) {
user_vec_f.push_back(rbwf.shoot());
}
vec_float_count += number;}
{std::vector<double>& user_vec_d = col_vec_d->variable();
user_vec_d.clear();
unsigned int number = row_wise ? count%100 : (unsigned int)(10*rflat.shoot());
for(unsigned int i=0;i<number;i++) {
user_vec_d.push_back(rg.shoot());
}
vec_double_count += number;}
{user_vec_s.clear();
unsigned int number = row_wise ? count%5 : (unsigned int)(5*rflat.shoot());
for(unsigned int i=0;i<number;i++) {
if(!inlib::num2s(i,stmp)){}
user_vec_s.push_back(stmp);
}}
if(!ntu->add_row()) {
std::cout << "ntuple fill failed." << std::endl;
break;
}
}
if(verbose) {
std::cout << "vec_float_count " << vec_float_count << std::endl;
std::cout << "vec_double_count " << vec_double_count << std::endl;
}}
//////////////////////////////////////////////////////////
/// create a ntuple from a ntuple_booking object. ////////
//////////////////////////////////////////////////////////
{inlib::ntuple_booking nbk("rg_rbw_2","Randoms");
nbk.add_column<double>("rgauss");
nbk.add_column<float>("rbw");
nbk.add_column<std::string>("string");
//nbk.add_column<bool>("not_handled");
inlib::wroot::ntuple* ntu = new inlib::wroot::ntuple(rfile.dir(),nbk);
if(ntu->columns().size()) {
inlib::wroot::ntuple::column<double>* col_rgauss = ntu->find_column<double>("rgauss");
inlib::wroot::ntuple::column<float>* col_rbw = ntu->find_column<float>("rbw");
inlib::wroot::ntuple::column_string* col_str = ntu->find_column_string("string");
inlib::rbwf rbwf(0,1);
std::string stmp;
for(unsigned int count=0;count<1000;count++) {
if(!col_rgauss->fill(rg.shoot())) {
std::cout << "col_rgauss fill failed." << std::endl;
break;
}
if(!col_rbw->fill(rbwf.shoot())) {
std::cout << "col_rbw fill failed." << std::endl;
break;
}
if(!inlib::num2s(count,stmp)){}
if(!col_str->fill("str "+stmp)) {
std::cout << "col_str fill failed." << std::endl;
break;
}
if(!ntu->add_row()) {
std::cout << "ntuple fill failed." << std::endl;
break;
}
}
}}
//////////////////////////////////////////////////////////
/// consistency check : create an empty ntuple : /////////
//////////////////////////////////////////////////////////
{inlib::wroot::ntuple* ntu = new inlib::wroot::ntuple(rfile.dir(),"empty","empty");
ntu->create_column<int>("empty");}
//////////////////////////////////////////////////////////
/// write and close file : ///////////////////////////////
//////////////////////////////////////////////////////////
{unsigned int n;
if(!rfile.write(n)) {
std::cout << "file write failed." << std::endl;
}}
rfile.close();}
//////////////////////////////////////////////////////////
/// read the file : //////////////////////////////////////
//////////////////////////////////////////////////////////
#include "read_root.icc"
//////////////////////////////////////////////////////////
//////////////////////////////////////////////////////////
//////////////////////////////////////////////////////////
#ifdef INLIB_MEM
}inlib::mem::balance(std::cout);
#endif //INLIB_MEM
return EXIT_SUCCESS;
}
The rroot.cpp program
// Copyright (C) 2010, Guy Barrand. All rights reserved.
// See the file exlib.license for terms.
//exlib_build_use exlib inlib csz zlib
#ifdef INLIB_MEM
#include <inlib/mem>
#endif //INLIB_MEM
#include <inlib/args>
#include <inlib/fileis>
#include <inlib/rroot/file>
#include <inlib/rroot/rall>
#include <inlib/rroot/ntuple>
#include <inlib/ntuple_binding>
#ifdef EXLIB_DONT_HAVE_ZLIB
#else
#include <exlib/zlib>
#endif
#include <iostream>
#include <cstdlib>
int main(int argc,char** argv) {
#ifdef INLIB_MEM
inlib::mem::set_check_by_class(true);{
#endif //INLIB_MEM
inlib::args args(argc,argv);
std::string file;
if(!args.file(file)) {
std::cout << "give a root file." << std::endl;
return EXIT_FAILURE;
}
bool verbose = args.is_arg("-verbose");
bool ls = args.is_arg("-ls");
bool dump = args.is_arg("-dump");
{bool is;
inlib::file::is_root(file,is);
if(!is) {
std::cout << " file is not a root file." << std::endl;
return EXIT_FAILURE;
}}
inlib::rroot::file rfile(std::cout,file,verbose);
#ifdef EXLIB_DONT_HAVE_ZLIB
#else
rfile.add_unziper('Z',exlib::decompress_buffer);
#endif
if(ls) {
std::cout << "format version " << rfile.version() << std::endl;
}
const std::vector<inlib::rroot::key*>& keys = rfile.dir().keys();
inlib::rroot::read(std::cout,rfile,keys,true,ls,dump,0);
///////////////////////////////////////////////////////////////
/// if reading the wroot.root produced with wroot.cpp : ///////
///////////////////////////////////////////////////////////////
{inlib::rroot::TDirectory* dir = inlib::rroot::find_dir(rfile.dir(),"histo");
if(dir) {
{inlib::rroot::key* key = dir->find_key("rg");
if(key) {
inlib::histo::h1d* h = inlib::rroot::key_to_h1d(rfile,*key);
if(h) {
std::cout << "h1d : " << h->title()
<< ", all_entries " << h->all_entries()
<< ", entries " << h->entries()
<< ", mean " << h->mean() << ", rms " << h->rms()
<< std::endl;
delete h;
}
}}
{inlib::rroot::key* key = dir->find_key("rf");
if(key) {
inlib::histo::h1d* h = inlib::rroot::key_to_h1d(rfile,*key);
if(h) {
std::cout << "h1d : " << h->title()
<< ", all_entries " << h->all_entries()
<< ", entries " << h->entries()
<< ", mean " << h->mean() << ", rms " << h->rms()
<< std::endl;
delete h;
}
}}
{inlib::rroot::key* key = dir->find_key("rgbw");
if(key) {
inlib::histo::h2d* h = inlib::rroot::key_to_h2d(rfile,*key);
if(h) {
std::cout << "h2d : " << h->title()
<< ", all_entries " << h->all_entries()
<< ", entries " << h->entries()
<< ", mean_x " << h->mean_x() << ", rms_x " << h->rms_x()
<< ", mean_y " << h->mean_y() << ", rms_y " << h->rms_y()
<< std::endl;
delete h;
}
}}
{inlib::rroot::key* key = dir->find_key("prof");
if(key) {
inlib::histo::p1d* h = inlib::rroot::key_to_p1d(rfile,*key);
if(h) {
std::cout << "p1d : " << h->title()
<< ", all_entries " << h->all_entries()
<< ", entries " << h->entries()
<< ", mean " << h->mean() << ", rms " << h->rms()
<< std::endl;
delete h;
}
}}
{inlib::rroot::key* key = dir->find_key("prof2D");
if(key) {
inlib::histo::p2d* h = inlib::rroot::key_to_p2d(rfile,*key);
if(h) {
std::cout << "p2d : " << h->title()
<< ", all_entries " << h->all_entries()
<< ", entries " << h->entries()
<< ", mean_x " << h->mean_x() << ", rms_x " << h->rms_x()
<< ", mean_y " << h->mean_y() << ", rms_y " << h->rms_y()
<< std::endl;
delete h;
}
}}
{inlib::rroot::key* key = dir->find_key("rggbw");
if(key) {
inlib::histo::h3d* h = inlib::rroot::key_to_h3d(rfile,*key);
if(h) {
std::cout << "h3d : " << h->title()
<< ", all_entries " << h->all_entries()
<< ", entries " << h->entries()
<< ", mean_x " << h->mean_x() << ", rms_x " << h->rms_x()
<< ", mean_y " << h->mean_y() << ", rms_y " << h->rms_y()
<< ", mean_z " << h->mean_z() << ", rms_z " << h->rms_z()
<< std::endl;
delete h;
}
}}
delete dir;
}}
////////////////////////////////////////////////////////////////////////////////////////
// read an ntuple from inlib/examples/cpp/wroot.cpp, wroot_pntuple.cpp, pwroot.cpp : ///
////////////////////////////////////////////////////////////////////////////////////////
{inlib::rroot::key* key = rfile.dir().find_key("rg_rbw");
if(key) {
unsigned int sz;
char* buf = key->get_object_buffer(rfile,sz);
if(!buf) {
std::cout << "can't get data buffer for ntuple." << std::endl;
return EXIT_FAILURE;
}
inlib::rroot::buffer b(std::cout,rfile.byte_swap(),sz,buf,key->key_length(),verbose);
inlib::rroot::fac fac(std::cout);
inlib::rroot::tree tree(rfile,fac);
if(!tree.stream(b)) {
std::cout << "TTree streaming failed." << std::endl;
return EXIT_FAILURE;
}
tree.dump(std::cout,""," ");
//inlib::uint64 entries = tree.entries();
{for(inlib::uint32 i=0;i<5;i++){
if(!tree.show(std::cout,i)) {
std::cout << "show failed for entry " << i << std::endl;
return EXIT_FAILURE;
}
}}
{inlib::uint64 entries = tree.entries();
for(inlib::uint64 i=inlib::mx<inlib::int64>(5,entries-5);i<entries;i++){
if(!tree.show(std::cout,(inlib::uint32)i)) {
std::cout << "show failed for entry " << i << std::endl;
return EXIT_FAILURE;
}
}}
// read with the flat ntuple API :
{inlib::rroot::ntuple ntu(tree); //use the flat ntuple API.
double user_rgauss;
std::string user_string;
inlib::ntuple_binding nbd;
nbd.add_column("rgauss",user_rgauss);
nbd.add_column("string",user_string);
if(!ntu.initialize(std::cout,nbd)) {
std::cout << "can't initialize ntuple with ntuple_binding." << std::endl;
return EXIT_FAILURE;
}
inlib::histo::h1d hg("rgauss",100,-5,5);
ntu.start();
unsigned int count = 0;
while(ntu.next()){
if(!ntu.get_row()) {
std::cout << "get_row() failed." << std::endl;
return EXIT_FAILURE;
}
hg.fill(user_rgauss);
if(count<5) std::cout << "user_string " << user_string << std::endl;
count++;
}
std::cout << "ntuple_binding(rgauss) : " << hg.mean() << " " << hg.rms() << std::endl;}
}}
///////////////////////////////////////////////////////////////
/// if reading the pawdemo.root : /////////////////////////////
///////////////////////////////////////////////////////////////
{inlib::rroot::key* key = rfile.dir().find_key("h10");
if(key) {
inlib::histo::h1d* h = inlib::rroot::key_to_h1d(rfile,*key);
if(h) {
std::cout << "h1d : h10"
<< ", all_entries " << h->all_entries()
<< ", entries " << h->entries()
<< ", mean " << h->mean() << ", rms " << h->rms()
<< std::endl;
delete h;
}
}}
/////////////////////////////////////////////////////////////////////
/// if reading the prof.root produced with croot_TProfile.cpp : /////
/////////////////////////////////////////////////////////////////////
{inlib::rroot::key* key = rfile.dir().find_key("prof");
if(key) {
inlib::histo::p1d* h = inlib::rroot::key_to_p1d(rfile,*key);
if(h) {
std::cout << "p1d : prof"
<< ", all_entries " << h->all_entries()
<< ", entries " << h->entries()
<< ", mean " << h->mean() << ", rms " << h->rms()
<< std::endl;
delete h;
}
}}
{inlib::rroot::key* key = rfile.dir().find_key("prof2D");
if(key) {
inlib::histo::p2d* h = inlib::rroot::key_to_p2d(rfile,*key);
if(h) {
std::cout << "p2d : prof"
<< ", all_entries " << h->all_entries()
<< ", entries " << h->entries()
<< ", mean_x " << h->mean_x() << ", rms_x " << h->rms_x()
<< ", mean_y " << h->mean_y() << ", rms_y " << h->rms_y()
<< std::endl;
delete h;
}
}}
#ifdef INLIB_MEM
}inlib::mem::balance(std::cout);
#endif //INLIB_MEM
return EXIT_SUCCESS;
}